Difference between revisions of "Robertisla"
Robertisla (talk | contribs) |
Robertisla (talk | contribs) |
||
| Line 615: | Line 615: | ||
|E Ion (au) || -264.58913 || -264.58913 || -303.68736 || -457.945733 | |E Ion (au) || -264.58913 || -264.58913 || -303.68736 || -457.945733 | ||
|- | |- | ||
| − | |E (HMim-Cl) (au) || -722.687898 || -722. | + | |E (HMim-Cl) (au) || -722.687898 || -722.66620 || -761.77953 || |
|- | |- | ||
| − | |Association energy ΔE (au) || -0.15301 || -0. | + | |Association energy ΔE (au) || -0.15301 || -0.13130 || -0.146437 || |
|- | |- | ||
| − | |Association energy (kJ/mol) || -401. | + | |Association energy (kJ/mol) || -401.7 || -344.7 || -384.5 || |
|} | |} | ||
'''Provide the relative energy of the two isomers (a) and (b)'''<br> | '''Provide the relative energy of the two isomers (a) and (b)'''<br> | ||
| − | ΔE = E<sub>b</sub> - E<sub>a</sub> = (-722.6662) - (-722.687898) = 0.021698 --> ΔE = | + | ΔE = E<sub>b</sub> - E<sub>a</sub> = (-722.6662) - (-722.687898) = 0.021698 --> ΔE = 57.0 kJ/mol <br> |
<br> | <br> | ||
This illustrates that the HMim-Cl B conformer is higher in energy than the HMim-Cl A conformer. This proves that the HMim-Cl A conformer has lower energy and is therefore a more stable conformer than B with greater polarity of bonds resulting in a stronger more favorable conformation. | This illustrates that the HMim-Cl B conformer is higher in energy than the HMim-Cl A conformer. This proves that the HMim-Cl A conformer has lower energy and is therefore a more stable conformer than B with greater polarity of bonds resulting in a stronger more favorable conformation. | ||
| Line 629: | Line 629: | ||
The HMim-Cl A conformer is the most stable as it has the largest association energy indicating that the molecule has a stronger more stable bond and it is less favourable to dissociate in comparison to the other confomers.<br> | The HMim-Cl A conformer is the most stable as it has the largest association energy indicating that the molecule has a stronger more stable bond and it is less favourable to dissociate in comparison to the other confomers.<br> | ||
<br> | <br> | ||
| − | The association energy of the HMim-Cl C conformer (-384. | + | The association energy of the HMim-Cl C conformer (-384.5 kJ/mol) lies in between the association energies of the HMim-Cl conformers A and B. The HMim-Cl B conformer has an association energy of -344.7 kJ/mol. Due to the lower association energy we can determine that the molecule is the least stable and the H-bonding is weaker. Conversely the HMim-Cl A conformer has a association energy of -401.7 kJ/mol this is larger than the HMim-Cl C conformer showing that although the H-bonding of this conformer is stronger than the HMim-Cl B |
conformer the H-bonding in the HMim-Cl A conformer is the strongest. | conformer the H-bonding in the HMim-Cl A conformer is the strongest. | ||
Revision as of 03:12, 15 May 2026
Contents
- 1 Lab1 Marking
- 2 BH3 Molecule
- 3 BH3NH3 Molecule
- 4 Me3NH-CL Molecule
- 5 1-methyl-imidazolium chloride A
- 6 1-methyl-imidazolium chloride B
- 7 1-methyl-imidazolium chloride C
- 8 Ionic liquid ion pair (1-methyl-imidazolium chloride (HMim-Cl) A, B, and C)
- 8.1 Optimised H--Cl bond lengths
- 8.2 Associated energies from 1-methyl-imidazolium chloride (HMim-Cl) A, B, and C
- 8.2.1 Optimised Ion and Jmol rotatable molecule for 1-methyl-imidazolium chloride ion A & B
- 8.2.2 Calculation data
- 8.2.3 Item Table
- 8.2.4 Low frequencies data
- 8.2.5 Optimised Chlorine atom and Jmol rotatable molecule
- 8.2.6 Calculation data
- 8.2.7 Item Table
- 8.2.8 Low frequencies data
- 8.2.9 Optimised Ion and rotatable molecule for 1-methyl-imidazolium chloride ion C
- 8.2.10 Calculation data
- 8.2.11 Item Table
- 8.2.12 Low frequencies data
- 8.2.13 Table of Association energies
- 8.3 PES rigid scan graph comparing Me3NH-Cl and (HMim)-Cl A
- 9 NH3 Molecule
Lab1 Marking
You did a great job especially with the formatting. However, you have tabulated wrong charges for N and F. If you have any queries, please contact Prof. Hunt.
BH3 Molecule
Calculation data
| name of submitted log file | IR_BH3_OPTF_POP.LOG |
| molecule | NH3 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -26.615112 |
| RMS gradient | 0.003053 |
| point group | D3h |
Item Table
Item Value Threshold Converged? Maximum Force 0.000004 0.000015 YES RMS Force 0.000003 0.000010 YES Maximum Displacement 0.000017 0.000060 YES RMS Displacement 0.000011 0.000040 YES
Low frequencies data
| Low frequencies | -12 | -12 | -7 | 0 | 0 | 0 |
| Low frequencies | 1163 | 1213 | 1213 |
Optimised molecule image
Jmol rotateable molecule
logfile: Media:IR BH3 OPTF POP.LOG
optimised BH molecule |
Important geometric parameters
Optimised bond distance and angle for BH3
r(B-H)=1.190Â
θ(H-B-H)=120.0°
Vibrational data
| Mode | 1 | 2 | 3 | 4 | 5 | 6 |
| Wavenumber (cm-1) | 1163 | 1213 | 1213 | 2583 | 2716 | 2716 |
| Symmetry | A2" | E' | E' | A1' | E' | E' |
| Intensity (arbitrary) | 93 | 14 | 14 | 0 | 126 | 126 |
BH3NH3 Molecule
Calculation data
| name of submitted log file | IR_NH3BH3_OPT.LOG |
| molecule | BH3NH3 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -83.22489 |
| RMS gradient | 0.000001 |
| point group | C1 |
Item Table
Item Value Threshold Converged? Maximum Force 0.000001 0.000015 YES RMS Force 0.000001 0.000010 YES Maximum Displacement 0.000043 0.000060 YES RMS Displacement 0.000019 0.000040 YES
Low frequencies data
| Low frequencies | -5 | -3 | 0 | 0 | 0 | 1 |
| Low frequencies | 263 | 633 | 638 |
Optimised molecule image
Jmol rotateable molecule
logfile: Media:IR_NH3BH3_OPT.LOG
optimised BHNH molecule |
Important geometric parameters
Optimised bond distances for BH3 NH3
r(B-H)=1.210Â
r(N-H)=1.018Â
r(N-B)=1.668Â
Optimised bond angles for BH3 NH3
θ(H-B-H)=114.0°
θ(H-N-H)=108.0°
θ(H-N-B)=111.0°
θ(H-B-N)=105.0°
Vibrational data
| Mode | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 |
| Wavenumber (cm-1) | 263 | 633 | 638 | 638 | 1069 | 1069 | 1196 | 1204 | 1204 | 1329 | 1676 | 1676 | 2472 | 2532 | 2532 | 3464 | 3581 | 3581 |
| Symmetry | A | A | A | A | A | A | A | A | A | A | A | A | A | A | A | A | A | A |
| Intensity (arbitrary) | 0 | 14 | 4 | 4 | 41 | 41 | 109 | 3 | 3 | 114 | 28 | 28 | 67 | 231 | 231 | 3 | 28 | 28 |
Total energies
E (NH3) = -56.557769
E (BH3) = -26.615112
E (BH3NH3) = -83.22489
Association Energy
ΔE= E(NH3BH3)-[E(NH3)+E(BH3)]
ΔE= -0.051989 Au
ΔE= -136.5 kJ/mol
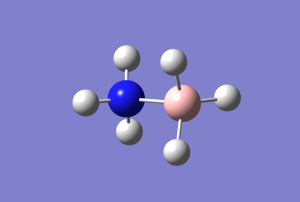
Me3NH-CL Molecule
Calculation data
| name of submitted log file | IR_ME3NH-CL_OPT2.LOG |
| molecule | Me3NH-Cl |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -26.615112 |
| RMS gradient | 0.003053 |
| point group | D3H |
Item Table
Item Value Threshold Converged? Maximum Force 0.000019 0.000450 YES RMS Force 0.000005 0.000300 YES Maximum Displacement 0.001834 0.001800 NO RMS Displacement 0.000343 0.001200 YES
Low frequencies data
| Low frequencies | -14 | -2 | 0 | 0 | 0 | 2 |
| Low frequencies | 56 | 56 | 190 |
Optimised molecule image
Jmol rotateable molecule
logfile: Media:IR_ME3NH-CL_OPT2.LOG
optimised MeNH-Cl molecule |
Important geometric parameters
Optimised bond lengths for Me3NH-Cl
r(N-H)= 1.164
r(H-Cl)= 1.738
r(N-Cl)= 2.902
Rigid Scan for Me3NH-Cl
PES rigid scan raw data plot for Me3NH-Cl
PES rigid scan raw data table for Me3NH-Cl
| Scan Coordinate  | Total Energy (Hartrees) | Relative Total Energy kJ mol -1 |
| 0.800 | -632.066250 | 229.9 |
| 0.900 | -632.122419 | 82.4 |
| 1.000 | -632.146048 | 20.4 |
| 1.100 | -632.153507 | 0.8 |
| 1.200 | -632.153801 | 0 |
| 1.300 | -632.151576 | 5.8 |
| 1.400 | -632.149019 | 12.6 |
| 1.500 | -632.147017 | 17.8 |
| 1.600 | -632.145741 | 21.2 |
| 1.700 | -632.144784 | 23.7 |
| 1.800 | -632.142934 | 28.5 |
| 1.900 | -632.137592 | 42.6 |
| 2.000 | -632.123855 | 78.6 |
| 2.100 | -632.093222 | 159.1 |
HMim-CL PES Plot for Me3NH-Cl
Plot of Total relative energy (kJ/mol) vs Scan coordinate (Â)
This graph shows that when the N-Cl distance is set at 3.2 Â a broad minima occurs, the most stable state is an ion-pair Me3NH+ --- Cl- and as the proton is pushed to the Cl forming a neutral pair Me3N --- HCl the energy goes up, and no stable minima forms, rather there is a "shelf" in the PES. The ion-pair forms a doubly ionic H-bond between the Me3NH+ and Cl-, the neutral-pair forms a normal H-bond between the Me3N and HCl.
1-methyl-imidazolium chloride A
Calculation data
| name of submitted log file | IR_IMIDA_A_OPT.LOG |
| molecule | 1-methyl-imidazolium chloride A |
| method | RB3LYP |
| basis set | 3-21G(d,p) |
| final energy | -722.687898 |
| RMS gradient | 1.96e-07 |
| point group | C1 |
Item Table
Item Value Threshold Converged? Maximum Force 0.000000 0.000015 YES RMS Force 0.000000 0.000010 YES Maximum Displacement 0.000004 0.000060 YES RMS Displacement 0.000001 0.000040 YES
Low frequencies data
| Low frequencies | -5 | -3 | 0 | 0 | 0 | 3 |
| Low frequencies | 36 | 64 | 81 |
Optimised molecule image
logfile: Media:IR IMIDA A OPT.LOG
Important Geometric parameters
Optimised bond lengths
r(H-N)(3-5)= 1.1776 Â
r(H-Cl)(3-14)= 1.719 Â
r(N-Cl)(5-14)= 2.891 Â
r(C-H)(1-2)= 1.075 Â
r(C-H)(12-13)= 1.074 Â
r(N-C)(5-12)= 1.390 Â
r(C-N)(5-1)= 1.332 Â
Scan for 1-methyl-imidazolium chloride A
Raw data
Data Table
| Scan Coordinate  | Total Energy (Hartrees) | Relative Total Energy kJ mol -1 |
| 0.800 | -722.595812 | 219.4 |
| 0.900 | -722.650398 | 76.1 |
| 1.000 | -722.672765 | 17.3 |
| 1.100 | -722.679373 | 0 |
| 1.200 | -722.679291 | 0.2 |
| 1.300 | -722.677213 | 5.7 |
| 1.400 | -722.675324 | 10.6 |
| 1.500 | -722.674443 | 12.9 |
| 1.600 | -722.674629 | 12.5 |
| 1.700 | -722.675355 | 10.6 |
| 1.800 | -722.675349 | 10.6 |
| 1.900 | -722.672137 | 19.0 |
| 2.000 | -722.661301 | 47.4 |
| 2.100 | -722.635447 | 115.3 |
Formal Scan Graph for 1-methyl-imidazolium chloride A
1-methyl-imidazolium chloride B
Calculation data
| name of submitted log file | IR_IMIDA_B_OPT.LOG |
| molecule | 1-methyl-imidazolium chloride B |
| method | RB3LYP |
| basis set | 3-21G(d,p) |
| final energy | -722.66620 |
| RMS gradient | 3.291e-06 |
| point group | C1 |
Item Table
Item Value Threshold Converged? Maximum Force 0.000006 0.000450 YES RMS Force 0.000002 0.000300 YES Maximum Displacement 0.000443 0.001800 YES RMS Displacement 0.000078 0.001200 YES
Low frequencies data
| Low frequencies | -5 | -3 | 0 | 0 | 0 | 2 |
| Low frequencies | 45 | 162 | 199 |
Optimised molecule image and Jmol rotateable molecule
logfile: Media:IR IMIDA B OPT.LOG
Important Geometric parameters
Optimised bond lengths
r(H-N)(7-3)= 1.1044 Â
r(H-Cl)(7-14)= 2.134 Â
r(H-Cl)(13-14)= 2.277 Â
r(N-Cl)(8-14)= 3.650 Â
r(C-H)(3-7)= 1.1045 Â
r(C-H)(10-13)= 1.106 Â
r(N-C)(8-10)= 1.496 Â
r(C-N)(3-8)= 1.402 Â
1-methyl-imidazolium chloride C
Calculation data
| name of submitted log file | IR_IMIDA_C_OPT.LOG |
| molecule | 1-methyl-imidazolium chloride C |
| method | RB3LYP |
| basis set | 3-21G(d,p) |
| final energy | -761.77953 |
| RMS gradient | 2.192e-06 |
| point group | C1 |
Item Table
Item Value Threshold Converged? Maximum Force 0.000006 0.000450 YES RMS Force 0.000002 0.000300 YES Maximum Displacement 0.000110 0.001800 YES RMS Displacement 0.000036 0.001200 YES
Low frequencies data
| Low frequencies | -4 | -3 | 0 | 0 | 0 | 2 |
| Low frequencies | 52 | 103 | 107 |
Optimised molecule image
logfile: Media:IR IMIDA C OPT.LOG
Important Geometric parameters
Optimised bond lengths
r(H-C)(4-1)= 1.118 Â
r(H-Cl)(4-17)= 2.030 Â
r(H-Cl)(11-17)= 2.417 Â
r(N-Cl)(7-17)= 3.666 Â
r(C-H)(9-11)= 1.101 Â
r(N-C)(7-9)= 1.490 Â
r(C-N)(1-7)= 1.348 Â
Ionic liquid ion pair (1-methyl-imidazolium chloride (HMim-Cl) A, B, and C)
Optimised H--Cl bond lengths
| Molecule Name | HMim-Cl A | HMim-Cl B | HMim-Cl C | Me3NH-Cl |
| Bond distance (Â) | 1.719 Â | 2.134 Â | 2.030 Â | 1.738 |
| Additional H--Cl bond | N/A | 2.277 Â | N/A | N/A |
How do the H-bonds of the H--Cl of Me3NH and HMim compare and how do the H-bonds of the N-H and C-H H-bonds compare??
The H--Cl bonds of the Me3NH-Cl molecule (1.738 Â) is very similar to the H--Cl of the HMim-Cl A (1.719 Â), whereas the H--Cl bonds of HMim-Cl B (2.134 Â, 2.277 Â) and HMim-Cl C (2.030 Â)are larger. This is because the hydrogen of the Me3NH-Cl and HMim-Cl A is bonded to a nitrogen which leads to a more polar bond. The effects of this polarity cause stronger dipole dipole attraction leaving the hydrogen with a larger delta + which produces a greater attraction between the hydrogen and the negative chlorine. Additionally, the HMim-Cl B molecule shares the H--Cl bonding between two bonds therefore leading to a more delocalized interaction and the longest H--Cl bonds. This polarity also results in a Me3NH-Cl
N-H bond length of 1.164 Â and a HMim-Cl A N-H bond length of 1.178 Â. Both these bonds end up being longer than the typical C-H bond because, although in general the N-H bond is more polar and leads to a stronger bond, the additional interaction with chlorine results in the sharing of the hydrogen lengthing the N-H. This is in comparison to the C-H bonds found for each HMim-Cl molecle to range between 1.070 - 1.100 Â. The N-H bond in the HMim-Cl B molecule is opposite from the site of the chlorine interaction therefore it has a shorter bond length of 1.014 Â as the hydrogen is not being shared between the chlorine atom and the nitorgen.
Are these distances representative of a H-bond and will the ionic nature of the ions effect a distance based assessment of H-bonding?
van der Waals radius for chlorine = 1.75 Â
van der Waals radius for hydrogen = 1.20 Â
Sum of van der Waals radius = 1.75 + 1.20 = 2.95 Â
Comparing the bonds we obtained to the standard sum of the van de Waals radii there is a noticeable difference in lengths, with the sum of van der Waals being between 0.673 Â to 1.231 Â longer than all of the Cl associated molecule. These result indicate that the ionic nature of these molecules does have an effect a distance based assessment of H-bonding with the attractive interactions influencing the bond length.
Associated energies from 1-methyl-imidazolium chloride (HMim-Cl) A, B, and C
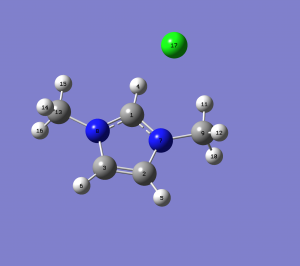
Optimised Ion and Jmol rotatable molecule for 1-methyl-imidazolium chloride ion A & B
 |
|
Calculation data
| name of submitted log file | IR_REATTEMPT_AB_ION.LOG |
| molecule | 1-methyl-imidazolium chloride ion A & B |
| method | RB3LYP |
| basis set | 3-21G |
| final energy | -264.58913 |
| RMS gradient | 1.0609e-05 |
| point group | C1 |
Item Table
Item Value Threshold Converged? Maximum Force 0.000134 0.000450 YES RMS Force 0.000028 0.000300 YES Maximum Displacement 0.000708 0.001800 YES RMS Displacement 0.000190 0.001200 YES
Low frequencies data
| Low frequencies | -5 | -4 | 0 | 0 | 0 | 4 |
| Low frequencies | 91 | 155 | 336 |
Optimised Chlorine atom and Jmol rotatable molecule
 |
|
Calculation data
| name of submitted log file | IR_CHLORINE_ATOM_OPT.LOG |
| molecule | Chlorine atom |
| method | UB3LYP |
| basis set | 3-21G |
| final energy | -457.94573 |
| RMS gradient | 0 |
| point group | OH |
Item Table
Item Value Threshold Converged? Maximum Force 0.000134 0.000450 YES RMS Force 0.000028 0.000300 YES Maximum Displacement 0.000708 0.001800 YES RMS Displacement 0.000190 0.001200 YES
Low frequencies data
| Low frequencies | 0 | 0 | 0 |
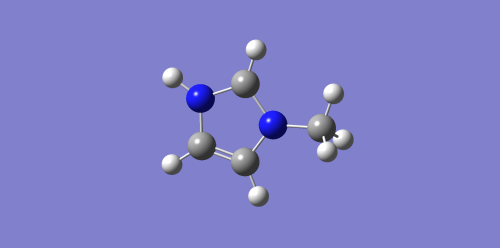
Optimised Ion and rotatable molecule for 1-methyl-imidazolium chloride ion C
 |
|
logfile: Media:IR_IMIDA_C_OPT_ION.LOG
Calculation data
| name of submitted log file | IR_IMIDA_C_OPT_ION.LOG |
| molecule | 1-methyl-imidazolium chloride ion C |
| method | RB3LYP |
| basis set | 3-21G |
| final energy | -303.68736 |
| RMS gradient | 2.4257e-05 |
| point group | C1 |
Item Table
Item Value Threshold Converged? Maximum Force 0.000134 0.000450 YES RMS Force 0.000028 0.000300 YES Maximum Displacement 0.000708 0.001800 YES RMS Displacement 0.000190 0.001200 YES
Low frequencies data
| Low frequencies | -870 | -200 | -191 | -77 | -53 | 0 |
| Low frequencies | 0 | 0 | 1 |
I theorize that the low frequencies have not meet the assumed range as the molecule that was optimized was the original optimized 1-methyl-imidazolium chloride (HMim-CL) C molecule that had it's chlorine atom removed and was optimized from there.
Table of Association energies
| Molecule | (HMim-Cl) A | (HMim-Cl) B | (HMim-Cl) C | Chloride anion |
| E Ion (au) | -264.58913 | -264.58913 | -303.68736 | -457.945733 |
| E (HMim-Cl) (au) | -722.687898 | -722.66620 | -761.77953 | |
| Association energy ΔE (au) | -0.15301 | -0.13130 | -0.146437 | |
| Association energy (kJ/mol) | -401.7 | -344.7 | -384.5 |
Provide the relative energy of the two isomers (a) and (b)
ΔE = Eb - Ea = (-722.6662) - (-722.687898) = 0.021698 --> ΔE = 57.0 kJ/mol
This illustrates that the HMim-Cl B conformer is higher in energy than the HMim-Cl A conformer. This proves that the HMim-Cl A conformer has lower energy and is therefore a more stable conformer than B with greater polarity of bonds resulting in a stronger more favorable conformation.
Rationalise why one conformer is less stable than the other and discuss the dissociation energy of (c) relative to (a) and (b). What does the comparison tell us about the H-bonding?
The HMim-Cl A conformer is the most stable as it has the largest association energy indicating that the molecule has a stronger more stable bond and it is less favourable to dissociate in comparison to the other confomers.
The association energy of the HMim-Cl C conformer (-384.5 kJ/mol) lies in between the association energies of the HMim-Cl conformers A and B. The HMim-Cl B conformer has an association energy of -344.7 kJ/mol. Due to the lower association energy we can determine that the molecule is the least stable and the H-bonding is weaker. Conversely the HMim-Cl A conformer has a association energy of -401.7 kJ/mol this is larger than the HMim-Cl C conformer showing that although the H-bonding of this conformer is stronger than the HMim-Cl B
conformer the H-bonding in the HMim-Cl A conformer is the strongest.
PES rigid scan graph comparing Me3NH-Cl and (HMim)-Cl A
Discuss your HMim-Cl PES plot, compare and contrast your results for the Me3NH-Cl and HMim-Cl PES.
The HMim-Cl A conformer has a minimum energy point at 1.1 Â while the Me3NH-Cl has a minimum energy point at 1.2Â demonstrating that both molecules most stable conformation lay at a similar distance under rigid constraints.
As the proton moves from the HMim / Me3N over to the chlorine the energy increases creating an shelf before the H bonds to the Cl forming the neutral-pair for both e.g. Me3N and HCl / HMim and HCl, and the energy rises rapidly.
This scan shows us that for both molecules the most stable, low-energy conformation involves the charged ion-pairs with the ionic nature playing a role in creating a stronger stabilization of each molecule lowering the overall energy.
NH3 Molecule
Calculation data
| name of submitted log file | IRobertson_nh3_optf_pop.log |
| molecule | NH3 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -56.557769 |
| RMS gradient | 1.53e-07 |
| point group | C3v |
Item Table
Item Value Threshold Converged?
Maximum Force 0.000000 0.000015 YES
RMS Force 0.000000 0.000010 YES
Maximum Displacement 0.000003 0.000060 YES
RMS Displacement 0.000001 0.000040 YES
Low frequencies data
| Low frequencies | -5.6864 | -3.6131 | -3.6124 | 0.0017 | 0.0048 | 0.0162 |
| Low frequencies | 1089.3674 | 1693.9284 | 1693.9284 |
Optimised molecule image
Jmol rotateable molecule
logfile: Media:IROBERTSON_NH3_OPTF_POP.LOG
optimised NH molecule |
Important geometric parameters
Optimised bond distance and angle for NH3
r(N-H)=1.02Â
θ(H-N-H)=106°
Vibrational data
| Mode | 1 | 2 | 3 | 4 | 5 | 6 |
| Wavenumber (cm-1) | 1089 | 1694 | 1694 | 3461 | 3590 | 3590 |
| Symmetry | A1 | E | E | A1 | E | E |
| Intensity (arbitrary) | 145 | 14 | 14 | 1 | 0 | 0 |
Optimised molecule image with charges
| Atoms | N | H |
| Charge | -1.13 | 0.38 |
N2F2 Molecule
Calculation data
| name of submitted log file | IJR_N2F2_C2V_OPTF.LOG |
| molecule | N2F2 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -309.01241 |
| RMS gradient | 3.17e-07 |
| point group | C2V |
Item Table
Item Value Threshold Converged? Maximum Force 0.000000 0.000015 YES RMS Force 0.000000 0.000010 YES Maximum Displacement 0.000001 0.000060 YES RMS Displacement 0.000000 0.000040 YES
Optimised N2F2 molecule
Optimised N2F2 molecule image
Jmol rotatable molecule
logfile: Media:IJR_N2F2_C2V_OPTF.LOG
optimised NF molecule |
Important geometric parameters
Optimised bond distance and angle for N2F2
r(N-F)=1.39Â
r(N-N)=1.22Â
θ(F-N-N)=114°
θ(F-N-N-F)=0°
Vibrational Data and analysis
| Mode | 1 | 2 | 3 | 4 | 5 | 6 |
| Wavenumber (cm-1) | 348 | 561 | 772 | 949 | 987 | 1637 |
| Symmetry | A1 | A2 | B2 | A1 | B2 | A1 |
| Intensity (arbitrary) | 1 | 0 | 75 | 75 | 81 | 21 |
Which vibration is the asymmetric N-F stretch?
Vibration 3 is asymmetric.
What is the nature of the highest energy vibration?
It is an N-N stretching vibration
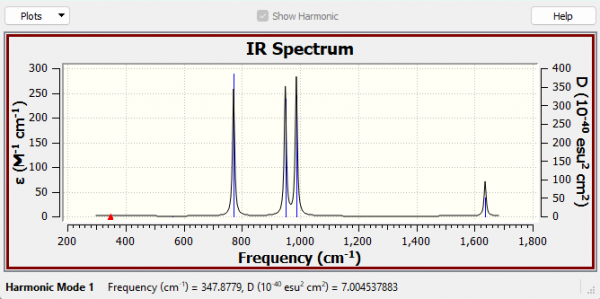
N2F2 Spectrum
IR Analysis
How many vibrations are expected from the 3N-6 rule?
We would expect there to be 6 vibrations (3(4) - 6 = 6)
Why are there only 4 peaks in the IR spectrum?
The A1 347.88 cm-1 and the A2 561.25 cm-1 modes have essentially no intensity, as both of these modes must have no dipole change. Therefore only the remaining 4 modes can be observed in the IR spectrum as peaks.
Low frequencies data
| Low frequencies | -0.0024 | -0.0016 | -0.0014 | 3.3364 | 4.3775 | 5.1348 |
| Low frequencies | 347.8779 | 561.2478 | 771.6088 |
Optimised molecule image with charges
| Atoms | N | F |
| Charge | 0.26 | -0.26 |
Molecular Orbital analysis
N2F2 MO 9
Which MOs are core orbital MOs?
The core MOs are the first four MOs 1-4 with low lying energy.