Difference between revisions of "KusabsOliv"
Kusabsoliv (talk | contribs) |
Kusabsoliv (talk | contribs) |
||
| Line 348: | Line 348: | ||
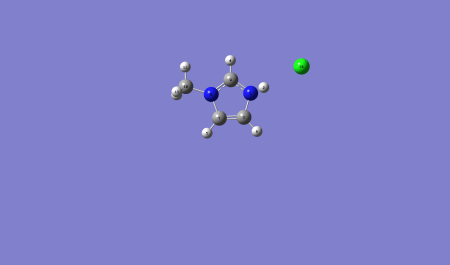
===Optimised Molecule=== | ===Optimised Molecule=== | ||
| − | [[File: | + | [[File:Kusabs_imida_B_optf_labelled.png|450px]] |
====Calculation Data==== | ====Calculation Data==== | ||
| Line 378: | Line 378: | ||
|} | |} | ||
| − | |||
logfile:[[Media:KUSABS_IMIDA_B_OPTF.LOG]] | logfile:[[Media:KUSABS_IMIDA_B_OPTF.LOG]] | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
Revision as of 05:21, 12 May 2026
Contents
- 1 Lab1 Marking
- 2 BH3 Molecule
- 3 NH3BH3 Molecule
- 4 Me3NH-Cl Molecule
- 5 Ionic Liquid pair 1-methyl Imadizolium chloride (HMim-Cl)
- 5.1 Molecule a)
- 5.2 Optimised molecule
- 5.3 Calculation data
- 5.4 Item Table
- 5.5 Low Frequencies
- 5.6 Jmol rotatable molecule
- 5.7 Important Geometric Parameters
- 5.8 HMim-Cl Molecule a) Scan
- 5.9 Hmim-Cl Formal Graph
- 5.10 1-methyl Imidazoliumm chloride B
- 5.11 Optimised Molecule
- 5.12 2-methyl Imidazolium chloride C
- 6 NH3 Molecule
- 7 N2F2 Molecule
Lab1 Marking
It's good that you have a working wiki. However, you have optimised N2F2 with a wrong symmetry, have reported wrong charges, and have missed to include the low frequencies, torsion angle, and NN distance. Don't forget to consider the accuracy to which you report your data the next time. If you have any queries, please contact Prof. Hunt.
BH3 Molecule
Optimized Molecule Image
"
calculation data
| molecule | BH3 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -26.615324 |
| RMS gradient | 0.000002 |
| point group | D3h |
Low frequencies
| Low frequencies | -11.6940 | -11.6861 | -6.5543 | 0.0007 | 0.0280 | 0.4289 |
| Low frequencies | 1162.9745 | 1213.1390 | 1213.1392 |
Item Table
Item Value Threshold Converged? Maximum Force 0.000004 0.000015 YES RMS Force 0.000003 0.000010 YES Maximum Displacement 0.000017 0.000060 YES RMS Displacement 0.000011 0.000040 YES
Jmol rotatable molecule
logfile:Media:KUSABSOLIV_BH3_OPTF_POP.LOG
optimised BHmolecule |
Important geometric parameters
optimized bond distance and angle for BH3
r(B-H)=1.19232Â
θ(H-B-H)=130°
Vibrational data
| mode | 1 | 2 | 2 | 4 | 5 | 6 |
| wavenumber(cm-1) | 1163 | 1213.14 | 1213.14 | 2582.58 | 2715.72 | 2715.72 |
| symmetry | A2" | E' | E' | A1' | E' | E' |
| intensity | 93 | 14 | 14 | 0 | 126 | 126 |
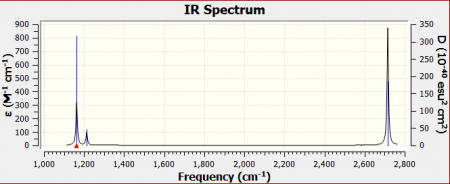
IR Spectrum
NH3BH3 Molecule
Optimized Molecule Image
calculation data
| molecule | NH3BH3 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -83.224689 |
| RMS gradient | 0.000001 |
| point group | C1 |
Item Table
Item Value Threshold Converged? Maximum Force 0.000001 0.000015 YES RMS Force 0.000001 0.000010 YES Maximum Displacement 0.000043 0.000060 YES RMS Displacement 0.000019 0.000040 YES
Low frequencies
| Low frequencies | -5.0440 | -2.8838 | 0.0010 | 0.0012 | 0.0014 | 0.6125 |
| Low frequencies | 263.3825 | 632.9842 | 638.4293 |
Jmol rotatable molecule
logfile:Media:OK_NH3BH3_OPTF_POP.LOG
optimised NHmolecule |
Important geometric parameters
optimized bond distance and angle for NH3BH3br
r(N-H)=1.01847Â
r(B-H)=1.20977Â
r=(B-N)=1.66770Â
θ(H-N-H)=108°
θ(H-B-H)=114°
θ(H-N-B)=105°
θ(H-B-N)=1118°
Vibrational data
| mode | 1 | 2 | 2 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 |
| wavenumber(cm-1) | 263 | 633 | 638 | 638 | 1069 | 1069 | 1196 | 1204 | 1204 | 1329 | 1676 | 1676 | 2472 | 2532 | 2532 | 3464 | 3581 | 3581 |
| symmetry | A | A | A | A | A | A | A | A | A | A | A | A | A | A | A | A | A | A |
| intensity | 0 | 14 | 4 | 4 | 41 | 41 | 109 | 3 | 3 | 114 | 28 | 28 | 67 | 231 | 231 | 3 | 28 | 28 |
IR spectrum
Association Energy
ΔE=E(NH3BH3)-[E(NH3)+E(BH3)]
=-0.05168 Hartrees
= -135.68584 kJ/mol
= 135.68584 kJ/mol (5 d.p)
Me3NH-Cl Molecule
Optimised Molecule
Calculation data
| molecule | Me3NH-Cl |
| method | RB3LYP |
| basis set | 3-21G |
| final energy | -632.16208 |
| RMS gradient | 5e-06 |
| point group | C1 |
Item Table
Item Value Threshold Converged?
Maximum Force 0.000008 0.000450 YES RMS Force 0.000003 0.000300 YES Maximum Displacement 0.000932 0.001800 YES RMS Displacement 0.000257 0.001200 YES
Low Frequencies
| Low frequencies | -13.6743 | -1.8717 | -0.0006 | 0.0026 | 0.0036 | 2.7014 |
| Low frequencies | 55.5069 | 56.3300 | 89.8235 |
Jmol Rotatable Image
logfile:Media:OK_ME3NHCL_OPTF_POP.LOG
optimised MeNHClmolecule |
Important Geometric Parameters
r(N-H)= 1.16Â
r(H-Cl)= 1.74 Â
r(N-Cl)= 2.90 Â
Me3HNCl Rigid Scan
Rigid Scan Raw Data Plot
PES Raw Data Table
| Scan Coordinate  | Total Energy (Hartrees) | Relative Total Energy (kJmol-1) |
| 0.8 | -632.0662517 | 229.8615 |
| 0.9 | -632.1224204 | 82.39055 |
| 1.0 | -632.1460690 | 20.30115 |
| 1.1 | -632.1535075 | 0.771372 |
| 1.2 | -632.1538013 | 0 |
| 1.3 | -632.1515761 | 5.842263 |
| 1.4 | -632.1490193 | 12.55514 |
| 1.5 | -632.1470175 | 17.81087 |
| 1.6 | -632.1457412 | 21.16179 |
| 1.7 | -632.1447843 | 23.67413 |
| 1.8 | -632.1429341 | 28.53183 |
| 1.9 | -632.1375920 | 42.55752 |
| 2.0 | -632.1238548 | 78.62454 |
| 2.1 | -632.0932227 | 159.0491 |
Me3NHCl Scan Formal Graph
Plot of Total Relative Energy (kJ/mol) vs. Scan Coordinate (Â)

The scan coordinate, which corresponds to the N-H bond length, starts at 0.8 Â and increasing in 0.1 Âincrements to 2.1 Â. When the N-Cl bond distance is set to 3.2 Â,the scan data plot shows that a minima occurs at 1.2Â. This displays an ion-pair Me3NH+ --- Cl-, and the minima proves that this is the most stable state. The graph captures the gradual shift of the proton between as the proton is pushed over to the Cl, where it forms a neutral pair Me3N---HCl. The energy goes up, with no stable minima formed, rather a 'shelf' appears in the PES. The ion-pair, forms a doubly ionic H-bond between the Me3NH+ and Cl-, the neutral pair forms a normal bond between Me3N and HCl.
Ionic Liquid pair 1-methyl Imadizolium chloride (HMim-Cl)
Molecule a)
Optimised molecule
Calculation data
| name of submitted log file | KUSABS_IMIDA_A_OPTF.LOG |
| molecule | HMim-Cl b) |
| method | RB3LYP |
| basis set | 3-21G |
| RMS gradient | 3.88e-07 |
| final energy | -722.6879 |
| point group | C1 |
Item Table
Item Value Threshold Converged? Maximum Force 0.000008 0.000450 YES RMS Force 0.000003 0.000300 YES Maximum Displacement 0.000932 0.001800 YES RMS Displacement 0.000257 0.001200 YES
Low Frequencies
| Low frequencies | -13.6743 | -1.8717 | -0.0006 | 0.0026 | 0.0036 | 2.7014 |
| Low frequencies | 55.5609 | 56.3300 | 189.8235 |
Jmol rotatable molecule
logfile:Media:OK_ME3NHCL_OPTF_POP.LOG
optimised HMim-Clmolecule |
Important Geometric Parameters
(9-7)r(N-H) =1.17767 Â
(9-14)r(N-Cl) = 2.89146 Â
(7-14)r(H-Cl) = 1.71933 Â
(1-4)r(C-H) = 1.07478 Â
(8-10)r(N-C) = 1.47703 Â
(2-5)r(C-H)=1.07442 Â
(2-8)r(C-N)=1.39999 Â
HMim-Cl Molecule a) Scan
HMim-Cl Scan Plot
PES Raw Data Table
| Scan Coordinate  | Total Energy (Hartrees) | Relative Total Energy (kJmol-1) |
| 0.8000000000 | -722.5958117770 | 219.390797 |
| 0.9000000000 | -722.6503978670 | 76.07501772 |
| 1.0000000000 | -722.6727650930 | 17.34986586 |
| 1.1000000000 | -722.6793733070 | 0 |
| 1.2000000000 | -722.6792906370 | 0.217050085 |
| 1.3000000000 | -722.6772128790 | 5.672203714 |
| 1.4000000000 | -722.6753243470 | 10.63054448 |
| 1.5000000000 | -722.6744432030 | 12.94398805 |
| 1.6000000000 | -722.6746286280 | 12.45715471 |
| 1.7000000000 | -722.6753546570 | 10.55096558 |
| 1.8000000000 | -722.6753493050 | 10.56501725 |
| 1.9000000000 | -722.6721372910 | 18.99816001 |
| 2.0000000000 | -722.6613012230 | 47.44825654 |
| 2.1000000000 | -722.6354468750 | 115.3288472 |
Hmim-Cl Formal Graph
Plot of Total Relative Energy (kJ/mol) vs. Scan Coordinate (Â)
1-methyl Imidazoliumm chloride B
Optimised Molecule
Calculation Data
| name of submitted log file | KUSABS_IMIDA_B_OPTF.LOG |
| molecule | 1-methyl Imidazoliumm chloride B |
| method | RB3LYP |
| basis set | 3-21G |
| Final energy | -722.666 |
| RMS gradient | 2.0678e-05 |
| Point group | C1 |
Item Table
Low Frequencies
| Low frequencies | -5.7142 | -3.2659 | -1.6460 | -0.0033 | -0.0016 | 0.0018 |
| Low frequencies | 45.3690 | 162.0925 | 198.8365 |
logfile:Media:KUSABS_IMIDA_B_OPTF.LOG
Important Geometric Parameters
(5-14) r(H-Cl)=2.13401 Â
(13-14)r(H-Cl)=2.27688 Â
(8-14) r(N-Cl)=3.64967 Â
(8-2) r(N-C)= 1.40258 Â
2-methyl Imidazolium chloride C
Jmol rotatable molecule
logfile:[[Media:KUSABS_IMIDA_C_OPTF2.LOG]
optimised HMim-Clmolecule |
NH3 Molecule
calculation data
| molecule | NH3 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -56.557769 |
| RMS gradient | 1.53e-07 |
| point group | C3v |
Item Table
Item Value Threshold Converged? Maximum Force 0.000000 0.000015 YES RMS Force 0.000000 0.000010 YES Maximum Displacement 0.000003 0.000060 YES RMS Displacement 0.000001 0.000040 YES
Optimized Molecule Image
Jmol rotatable molecule
logfile:Media:KUSABSOLIV_NH3_OPTF_POP.LOG
optimised NHmolecule |
Important geometric parameters
optimized bond distance and angle for NH3
r(N-H)=1.018Â
θ(H-N-H)=106°
Vibrational data
| mode | 1 | 2 | 2 | 4 | 5 | 6 |
| wavenumber(cm-1) | 1089 | 1694 | 1694 | 3461 | 3590 | 3590 |
| symmetry | A1 | E | E | A1 | E | E |
| intensity | 145 | 14 | 14 | 1 | 0 | 0 |
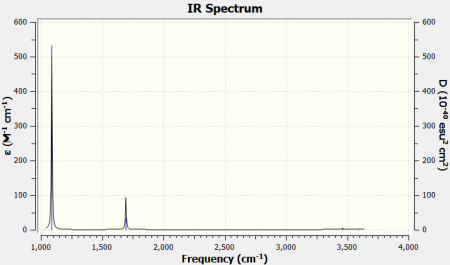
IR Spectrum
Charge Distribution Model
| Atom | Charge |
| Nitrogen | -1.13 |
| Hydrogen | 0.375 |
N2F2 Molecule
calculation data
| molecule | N2F2 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -309.01241 |
| RMS gradient | 3.685e-06 |
| point group | C2v |
Item Table
Item Value Threshold Converged? Maximum Force 0.000006 0.000015 YES RMS Force 0.000005 0.000010 YES Maximum Displacement 0.000024 0.000060 YES RMS Displacement 0.000017 0.000040 YES
Low frequencies
| Low frequencies | -0.0012 | -0.0012 | 0.0013 | 2.4177 | 4.2047 | 4.8781 |
| Low frequencies | 347.8622 | 561.2409 | 771.6170 |
Optimised Model Image
Jmol Rotatable Molecule
logfile:Media:KUSABSOLIV_N2F2_OPTF_POP.LOG
optimised NFmolecule |
Important geometric parameters
optimized bond distance and angle for N2F2
r(N-F)=1.39Â
θ(F-N-N)=114°
Vibrational data
| mode | 1 | 2 | 2 | 4 | 5 | 6 |
| wavenumber(cm-1) | 348 | 561 | 772 | 949 | 987 | 1637 |
| symmetry | A | A | B | A | B | A |
| intensity | 1 | 0 | 75 | 75 | 81 | 21 |
IR spectrum
Charge Distribution Model
| Atom | Charge |
| Nitrogen | -0.22 |
| Fluorine | 0.22 |
the molecule from the log file does not have bonds between the F and N atoms, what is going on here?
IR analysis
As there are 4 atoms in N2F2, 6 vibrations are expected from the 3N-6 rule. This matches to the 6 vibrations seen. There are four strong IR peaks shown at 772, 949, 989 and 1637. However, the peak at 348 does not show up as it is likely too low to detected, and the peak at 561 (A2, out of plane bending) is likely IR inactive as it does not change the molecules dipole. The vibration at 772cm-1, mode 3, is the asymmetric N-F stretch. The highest energy vibration at 1637cm-1 is the N=N double bond stretch.
Molecular Orbital Analysis
In N2F2, the core molecular orbitals are 1,2,3 and 4