Difference between revisions of "Youngeric"
| Line 220: | Line 220: | ||
'''Bonds''' | '''Bonds''' | ||
| − | + | No bonds between the F and N atoms since this is the most optimized. During this optimisation, the N=N bonds are closer and the N-F bond are further apart, virtually not showing the bonds. | |
'''IR analysis''' | '''IR analysis''' | ||
Revision as of 03:28, 2 April 2026
NH3 Molecule
calculation data
| name of submitted log file | EYOUNG_NH3_OPTF_POP.LOG |
| molecule | NH3 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -56.557769 |
| RMS gradient | 0.000000153 |
| point group | C3v |
Item Table
Item Value Threshold Converged? Maximum Force 0.000000 0.000015 YES RMS Force 0.000000 0.000010 YES Maximum Displacement 0.000003 0.000060 YES RMS Displacement 0.000001 0.000040 YES
Optimised molecule image
Jmol rotateable molecule
logfile: Media:EYOUNG_NH3_OPTF_POP.LOG
optimised NH molecule |
Important geometric parameters
Optimised bond distance and angle for NH3
r(N-H)=1.02Â
θ(H-N-H)=106°
Frequency
Low frequencies --- -5.6864 -3.6131 -3.6124 0.0017
0.0048 0.0162
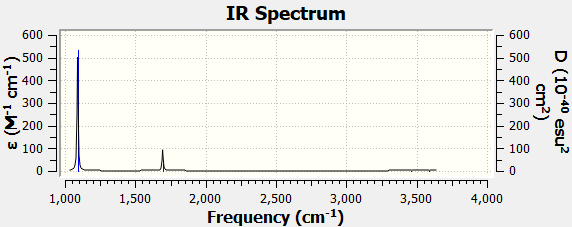
Low frequencies --- 1089.3674 1693.9284 1693.9284
| Mode | 1 | 2 | 3 | 4 | 5 | 6 |
|---|---|---|---|---|---|---|
| Wavenumber (cm-1) | 1089 | 1694 | 1694 | 3461 | 3590 | 3590 |
| Symmetry | A1 | E | E | A1 | E | E |
| Intensity (arbitrary units) | 145 | 14 | 14 | 1 | 0 | 0 |
Spectra
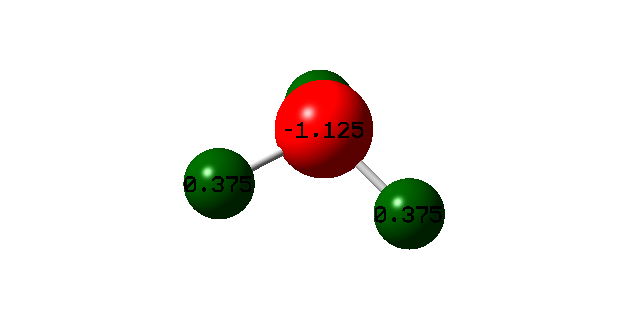
Atomic Charges
| Charges | |
| N-atom | -1.125 |
| H-atoms | 0.375 |
N2F2 Molecule
| name of submitted log file | EYOUNG_CIS(N2F2)_OPTF_POP |
| molecule | N2F2 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -308.98118 |
| RMS gradient | 0.0732 |
| point group | C2v |
Item Table
Item Value Threshold Converged? Maximum Force 0.000002 0.000015 YES RMS Force 0.000002 0.000010 YES Maximum Displacement 0.000010 0.000060 YES RMS Displacement 0.000006 0.000040 YES
Optimised molecule image
Jmol rotateable molecule
logfile: Media:EYOUNG_CISN2F2_OPTF_POP.LOG
optimised NF molecule |
Important geometric parameters
Optimised bond distance and angle for N2F2
r(N=N)=1.22Â
r(N-F)=1.39Â
θ(F-N-N)=114°
Frequency
Low frequencies --- -0.0007 -0.0005 -0.0003 3.2588 4.4213 5.1468 Low frequencies --- 347.8824 561.2505 771.6121
| Mode | 1 | 2 | 3 | 4 | 5 | 6 |
|---|---|---|---|---|---|---|
| Wavenumber (cm-1) | 348 | 561 | 772 | 949 | 987 | 1637 |
| Symmetry | A1 | A2 | B2 | A1 | B2 | A1 |
| Intensity (arbitrary units) | 1 | 0 | 75 | 75 | 81 | 21 |
Spectra
Atomic Charges
| Charges | |
|---|---|
| N-atom | 0.215 |
| F-atoms | -0.215 |
Questions
Bonds
No bonds between the F and N atoms since this is the most optimized. During this optimisation, the N=N bonds are closer and the N-F bond are further apart, virtually not showing the bonds.
IR analysis
- 3N-6 where N=the number of atoms, N=4 so 12-6=6, therefore we expect 6 modes which we have
- 4 peaks in IR due to some vibrations being inactive/weak specifically the twisting and wagging.
- Asymmetric N-F stretch is mode 3
- Highest energy vibration is mode 6, symmetric stretching of the N=N.
MO Analysis