Difference between revisions of "Wolfeka"
| (73 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
| + | ==to do== | ||
| + | *redo some of the accuracies | ||
| + | tabulate and compare all the H---Cl distances for all ion-pairs. | ||
| + | how do the H-bonds of Me3NH and HMim compare? | ||
| + | how do the H-bonds of the N-H and C-H H-bonds compare? | ||
| + | are these distances representative of a H-bond? Hint compare your distances to the sum of the van der waals radii of H and Cl. | ||
| + | will the ionic nature of the ions effect a distance based assessment of H-bonding? | ||
| + | provide a table and comparison of the association energy for a-c | ||
| + | make this explicit, so give the energy of each component in atomic units, compute the deltaE in au and then convert to kJ/mol | ||
| + | provide the relative energy of the two isomers (a) and (b), ask if you are not sure what this is. | ||
| + | rationalise why one conformer is less stable than the other. | ||
| + | discuss the dissociation energy of (c) relative to (a) and (b). What does the comparison tell us about the H-bonding? | ||
| + | scan the N-H coordinate towards Cl of (a) | ||
| + | take your optimised (a)structure, swap atoms to make N=1 and H=2, set the N-H distance to 0.8 and the N-Cl distance to 3.2, run the scan. This will take some time, start it early! | ||
| + | provide a snapshot from gaussview of the "raw" HMim-Cl PES plot | ||
| + | provide a chemical diagram of the two protonation states for HMim-Cl | ||
| + | graph the MeNH-Cl and HMim-Cl scans on a single PES plot | ||
| + | discuss your HMim-Cl PES plot, compare and contrast your results for the MeNH-Cl and HMim-Cl PES. | ||
| + | |||
| + | ==1-methyl-imidazolium chloride== | ||
| + | ===Calculation data=== | ||
| + | {| class="wikitable" | ||
| + | |name of submitted log file|| KATW_RINGA_OPTFREQ2.log | ||
| + | |- | ||
| + | |molecule|| 1-methyl-imidazolium chloride A | ||
| + | |- | ||
| + | |method|| RB3LYP | ||
| + | |- | ||
| + | |basis set|| 3-21G | ||
| + | |- | ||
| + | |final energy||-722.68731au | ||
| + | |- | ||
| + | |RMS gradient||5.198e-06au/bohr | ||
| + | |- | ||
| + | |point group|| C1 | ||
| + | |} | ||
| + | |||
| + | |||
| + | ===Item table=== | ||
| + | Optimizing item table | ||
| + | <pre> | ||
| + | Item Value Threshold Converged? | ||
| + | Maximum Force 0.000011 0.000450 YES | ||
| + | RMS Force 0.000003 0.000300 YES | ||
| + | Maximum Displacement 0.001137 0.001800 YES | ||
| + | RMS Displacement 0.000246 0.001200 YES | ||
| + | </pre> | ||
| + | |||
| + | Frequency item table | ||
| + | <pre> | ||
| + | Item Value Threshold Converged? | ||
| + | Maximum Force 0.000011 0.000450 YES | ||
| + | RMS Force 0.000003 0.000300 YES | ||
| + | Maximum Displacement 0.001332 0.001800 YES | ||
| + | RMS Displacement 0.000305 0.001200 YES | ||
| + | </pre> | ||
| + | |||
| + | |||
| + | ====Key bond distances and angles==== | ||
| + | *add another decimal point to each | ||
| + | {| class="wikitable" | ||
| + | |r(H-Cl) bond distance|| 1.719Â | ||
| + | |} | ||
| + | |||
| + | ===File history=== | ||
| + | [[Media:KATW RINGA OPTFREQ2.LOG]] | ||
| + | |||
| + | |||
| + | ===1-methyl-imidazolium chloride A frequencies=== | ||
| + | <pre> | ||
| + | Low frequencies --- -5.2962 -2.8134 -0.0015 0.0024 0.0026 2.7809 | ||
| + | Low frequencies --- 36.1361 64.4953 80.8261 | ||
| + | </pre> | ||
| + | The initial frequency scan resulted in a negative frequency vibration from the methyl group. The methyl group was manually rotated and the scan run again with reading internal force constants to yield better data. | ||
| + | |||
| + | ===Calculation data=== | ||
| + | {| class="wikitable" | ||
| + | |name of submitted log file|| KATW RINGB OPTFREQ2.LOG | ||
| + | |- | ||
| + | |molecule|| 1-methyl-imidazolium chloride B | ||
| + | |- | ||
| + | |method|| RB3LYP | ||
| + | |- | ||
| + | |basis set|| 3-21G | ||
| + | |- | ||
| + | |final energy||-722.6662au | ||
| + | |- | ||
| + | |RMS gradient||2.5631e-05au/bohr | ||
| + | |- | ||
| + | |point group|| C1 | ||
| + | |} | ||
| + | |||
| + | |||
| + | ===Item table=== | ||
| + | Optimizing item table | ||
| + | <pre> | ||
| + | Item Value Threshold Converged? | ||
| + | Maximum Force 0.000088 0.000450 YES | ||
| + | RMS Force 0.000020 0.000300 YES | ||
| + | Maximum Displacement 0.000693 0.001800 YES | ||
| + | RMS Displacement 0.000163 0.001200 YES | ||
| + | </pre> | ||
| + | |||
| + | Frequency item table | ||
| + | <pre> | ||
| + | Item Value Threshold Converged? | ||
| + | Maximum Force 0.000088 0.000450 YES | ||
| + | RMS Force 0.000020 0.000300 YES | ||
| + | Maximum Displacement 0.001343 0.001800 YES | ||
| + | RMS Displacement 0.000321 0.001200 YES | ||
| + | </pre> | ||
| + | |||
| + | |||
| + | ====Key bond distances and angles==== | ||
| + | *add another decimal point to each | ||
| + | {| class="wikitable" | ||
| + | |r(CH-Cl) bond distance|| 2.134Â | ||
| + | |- | ||
| + | |r(CH2H-Cl) bond distance|| 2.277Â | ||
| + | |} | ||
| + | |||
| + | ===File history=== | ||
| + | [[Media:KATW RINGB OPTFREQ2.LOG]] | ||
| + | |||
| + | |||
| + | ===1-methyl-imidazolium chloride B frequencies=== | ||
| + | <pre> | ||
| + | Low frequencies --- -6.2380 -3.1641 0.0011 0.0033 0.0040 1.3698 | ||
| + | Low frequencies --- 45.3380 162.1734 198.8517 | ||
| + | </pre> | ||
| + | |||
| + | |||
| + | ===Calculation data=== | ||
| + | {| class="wikitable" | ||
| + | |name of submitted log file|| KATW_RINGC_OPTFREQ.log | ||
| + | |- | ||
| + | |molecule|| 1-methyl-imidazolium chloride C | ||
| + | |- | ||
| + | |method|| RB3LYP | ||
| + | |- | ||
| + | |basis set|| 3-21G | ||
| + | |- | ||
| + | |final energy||-761.77953au | ||
| + | |- | ||
| + | |RMS gradient||8.305e-06au/bohr | ||
| + | |- | ||
| + | |point group|| C1 | ||
| + | |} | ||
| + | |||
| + | |||
| + | ===Item table=== | ||
| + | Optimizing item table | ||
| + | <pre> | ||
| + | Item Value Threshold Converged? | ||
| + | Maximum Force 0.000027 0.000450 YES | ||
| + | RMS Force 0.000005 0.000300 YES | ||
| + | Maximum Displacement 0.000555 0.001800 YES | ||
| + | RMS Displacement 0.000154 0.001200 YES | ||
| + | </pre> | ||
| + | |||
| + | Frequency item table | ||
| + | <pre> | ||
| + | Item Value Threshold Converged? | ||
| + | Maximum Force 0.000027 0.000450 YES | ||
| + | RMS Force 0.000005 0.000300 YES | ||
| + | Maximum Displacement 0.000581 0.001800 YES | ||
| + | RMS Displacement 0.000167 0.001200 YES | ||
| + | </pre> | ||
| + | |||
| + | |||
| + | ====Key bond distances and angles==== | ||
| + | {| class="wikitable" | ||
| + | |r(H-Cl) bond distance|| 2.030Â | ||
| + | |} | ||
| + | |||
| + | ===File history=== | ||
| + | [[Media:KATW RINGC OPTFREQ.LOG]] | ||
| + | |||
| + | |||
| + | ===1-methyl-imidazolium chloride C frequencies=== | ||
| + | <pre> | ||
| + | Low frequencies --- -3.7052 -3.0699 -0.0042 -0.0027 -0.0019 2.2969 | ||
| + | Low frequencies --- 52.1621 102.5835 107.1578 | ||
| + | </pre> | ||
| + | |||
| + | |||
| + | ===1-methyl-imidazolium chloride A scan of total energy=== | ||
| + | |||
| + | [[File:KATW SCANIMAGERINGA.PNG|350px]] | ||
| + | |||
| + | <b>Scan of Total Energy</b> | ||
| + | |||
| + | X-Axis: Scan Coordinate | ||
| + | |||
| + | Y-Axis: Total Energy (Hartree) | ||
| + | |||
| + | X Y | ||
| + | 0.8000000000 -722.5958115700 | ||
| + | 0.9000000000 -722.6503976760 | ||
| + | 1.0000000000 -722.6727649280 | ||
| + | 1.1000000000 -722.6793731750 | ||
| + | 1.2000000000 -722.6792905430 | ||
| + | 1.3000000000 -722.6772128260 | ||
| + | 1.4000000000 -722.6753243350 | ||
| + | 1.5000000000 -722.6744432350 | ||
| + | 1.6000000000 -722.6746287060 | ||
| + | 1.7000000000 -722.6753547910 | ||
| + | 1.8000000000 -722.6753495180 | ||
| + | 1.9000000000 -722.6721376400 | ||
| + | 2.0000000000 -722.6613018340 | ||
| + | 2.1000000000 -722.6354480070 | ||
| + | |||
| + | == Me<sub>3</sub>NH Cl molecule == | ||
| + | ===Calculation data=== | ||
| + | {| class="wikitable" | ||
| + | |name of submitted log file|| KATW_Me3NH-Clactual.log | ||
| + | |- | ||
| + | |molecule|| Me<sub>3</sub>NH Cl | ||
| + | |- | ||
| + | |method|| RB3LYP | ||
| + | |- | ||
| + | |basis set|| 6-31G(d,p) | ||
| + | |- | ||
| + | |final energy||-635.31113au | ||
| + | |- | ||
| + | |RMS gradient||4.0214e-05au/bohr | ||
| + | |- | ||
| + | |point group|| C1 | ||
| + | |} | ||
| + | |||
| + | The above point group was not set to be C3v, which it should be, so the program assigned it C1. | ||
| + | |||
| + | ===Item table=== | ||
| + | Optimizing item table | ||
| + | <pre> | ||
| + | Item Value Threshold Converged? | ||
| + | Maximum Force 0.000126 0.000450 YES | ||
| + | RMS Force 0.000032 0.000300 YES | ||
| + | Maximum Displacement 0.000835 0.001800 YES | ||
| + | RMS Displacement 0.000219 0.001200 YES | ||
| + | </pre> | ||
| + | |||
| + | Frequency item table | ||
| + | <pre> | ||
| + | Item Value Threshold Converged? | ||
| + | Maximum Force 0.000126 0.000450 YES | ||
| + | RMS Force 0.000032 0.000300 YES | ||
| + | Maximum Displacement 0.001285 0.001800 YES | ||
| + | RMS Displacement 0.000457 0.001200 YES | ||
| + | </pre> | ||
| + | |||
| + | ===Optimised molecule image=== | ||
| + | [[File:Me3ClNH KATW image.PNG|150px]] | ||
| + | <jmol><jmolApplet> | ||
| + | <title>3D Me3NH Cl molecule</title> | ||
| + | <color>#ff82a8</color> | ||
| + | <size>200</size> | ||
| + | <uploadedFileContents>KATW ME3NH-CLACTUAL.LOG</uploadedFileContents> | ||
| + | </jmolApplet></jmol> | ||
| + | |||
| + | ====Key bond distances and angles==== | ||
| + | *add another decimal point to each | ||
| + | {| class="wikitable" | ||
| + | |r(N-C) bond distance|| 1.48Â | ||
| + | |- | ||
| + | |r(N-H) bond distance|| 0.80Â | ||
| + | |- | ||
| + | |r(N-Cl) bond distance|| 3.20Â | ||
| + | |- | ||
| + | |r(C-H) bond distance|| 1.09Â | ||
| + | |- | ||
| + | | θ(C-N-C) bond angle|| 112° | ||
| + | |- | ||
| + | | θ(C-N-H) bond angle|| 106° | ||
| + | |- | ||
| + | | θ(H-C-H) bond angle|| 110° | ||
| + | |} | ||
| + | |||
| + | ===File history=== | ||
| + | [[Media:KATW ME3NH-CLACTUAL.LOG]] | ||
| + | |||
| + | |||
| + | |||
| + | ===NH<sub>3</sub>BH<sub>3</sub> frequencies=== | ||
| + | <pre> | ||
| + | Low frequencies --- -3.0077 -0.5001 -0.0059 -0.0053 -0.0047 6.2314 | ||
| + | Low frequencies --- 81.5384 81.6770 183.6441 | ||
| + | </pre> | ||
| + | |||
| + | ===NH<sub>3</sub>BH<sub>3</sub> scan of total energy=== | ||
| + | |||
| + | <b>Snapshot: </b> | ||
| + | |||
| + | X-Axis: Scan Coordinate <p> | ||
| + | </p>Y-Axis: Total Energy (Hartree) | ||
| + | X Y | ||
| + | 0.8000000000 -635.2162861320 | ||
| + | 0.9000000000 -635.2709452560 | ||
| + | 1.0000000000 -635.2930688430 | ||
| + | 1.1000000000 -635.2992906270 | ||
| + | 1.2000000000 -635.2988745370 | ||
| + | 1.3000000000 -635.2968157070 | ||
| + | 1.4000000000 -635.2956086330 | ||
| + | 1.5000000000 -635.2962523740 | ||
| + | 1.6000000000 -635.2987711990 | ||
| + | 1.7000000000 -635.3023546920 | ||
| + | 1.8000000000 -635.3052254510 | ||
| + | 1.9000000000 -635.3042681400 | ||
| + | 2.0000000000 -635.2943300920 | ||
| + | 2.1000000000 -635.2670421330 | ||
| + | |||
| + | <b>Scan of the potential energy surface original graph:</b> | ||
| + | |||
| + | [[File:KATW MeNHCl scan graph1.PNG|350px]] | ||
| + | |||
| + | ==Relative energies== | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |molecule||Energy (au)||Energy (kJ/mol)||3||4||5||6 | ||
| + | |- | ||
| + | |NH3||-56.557769||-148492||1213||2583||2716||2716 | ||
| + | |- | ||
| + | |BH3||-26.615324||-69879||E||A1||E||E | ||
| + | |- | ||
| + | |NH3BH3||-83.2246894||-218506||14||0||126||126 | ||
| + | |} | ||
| + | |||
| + | |||
| + | E(NH3BH3)= | ||
| + | |||
| + | == BH<sub>3</sub> molecule == | ||
| + | ===Calculation data=== | ||
| + | {| class="wikitable" | ||
| + | |name of submitted log file|| BH3_optimization_KATW.log | ||
| + | |- | ||
| + | |molecule|| BH<sub>3</sub> | ||
| + | |- | ||
| + | |method|| RB3LYP | ||
| + | |- | ||
| + | |basis set|| 6-31G(d,p) | ||
| + | |- | ||
| + | |final energy|| -26.615324au | ||
| + | |- | ||
| + | |RMS gradient||2.114e-06au/bohr | ||
| + | |- | ||
| + | |point group|| D3H | ||
| + | |} | ||
| + | |||
| + | ===Item table=== | ||
| + | <pre> | ||
| + | Item Value Threshold Converged? | ||
| + | Maximum Force 0.000004 0.000015 YES | ||
| + | RMS Force 0.000003 0.000010 YES | ||
| + | Maximum Displacement 0.000017 0.000060 YES | ||
| + | RMS Displacement 0.000011 0.000040 YES | ||
| + | </pre> | ||
| + | |||
| + | |||
| + | ===Optimised molecule image=== | ||
| + | [[File:KATW BH3 optimagemol.PNG|150px]] | ||
| + | <jmol><jmolApplet> | ||
| + | <title>3D BH<sub>3</sub> molecule</title> | ||
| + | <color>#ff82a8</color> | ||
| + | <size>200</size> | ||
| + | <uploadedFileContents>BH3 OPTIMIZATION KATW.LOG</uploadedFileContents> | ||
| + | </jmolApplet></jmol> | ||
| + | |||
| + | ====Key bond distances and angles==== | ||
| + | {| class="wikitable" | ||
| + | |r(B-H) bond distance|| 1.19Â | ||
| + | |- | ||
| + | | θ(H-B-H) bond angle|| 120° | ||
| + | |} | ||
| + | |||
| + | |||
| + | ===File history=== | ||
| + | [[Media:BH3 OPTIMIZATION KATW.LOG]] | ||
| + | |||
| + | |||
| + | |||
| + | ===BH<sub>3</sub> IR spectrum=== | ||
| + | [[File:KATW BH3 IR.PNG]] | ||
| + | |||
| + | ===BH<sub>3</sub> frequencies=== | ||
| + | <pre> | ||
| + | Low frequencies --- -11.6940 -11.6861 -6.5543 -0.0006 0.0280 0.4289 | ||
| + | Low frequencies --- 1162.9745 1213.1390 1213.1392 | ||
| + | </pre> | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |mode||1||2||3||4||5||6 | ||
| + | |- | ||
| + | |wavenumber (cm<sup>-1</sup>)||1163||1213||1213||2583||2716||2716 | ||
| + | |- | ||
| + | |symmetry||A2||E||E||A1||E||E | ||
| + | |- | ||
| + | |intensity||93||14||14||0||126||126 | ||
| + | |} | ||
| + | |||
| + | == NH<sub>3</sub>BH<sub>3</sub> molecule == | ||
| + | ===Calculation data=== | ||
| + | {| class="wikitable" | ||
| + | |name of submitted log file|| KATW_NH3BH3_opt.log | ||
| + | |- | ||
| + | |molecule|| NH<sub>3</sub>BH<sub>3</sub> | ||
| + | |- | ||
| + | |method|| RB3LYP | ||
| + | |- | ||
| + | |basis set|| 6-31G(d,p) | ||
| + | |- | ||
| + | |final energy|| -83.2246894au | ||
| + | |- | ||
| + | |RMS gradient||1.264e-06au/bohr | ||
| + | |- | ||
| + | |point group|| C1 | ||
| + | |} | ||
| + | |||
| + | ===Item table=== | ||
| + | <pre> | ||
| + | Item Value Threshold Converged? | ||
| + | Maximum Force 0.000002 0.000015 YES | ||
| + | RMS Force 0.000001 0.000010 YES | ||
| + | Maximum Displacement 0.000016 0.000060 YES | ||
| + | RMS Displacement 0.000007 0.000040 YES | ||
| + | </pre> | ||
| + | |||
| + | |||
| + | ===Optimised molecule image=== | ||
| + | [[File:KATW NH3BH3 image.PNG|150px]] | ||
| + | <jmol><jmolApplet> | ||
| + | <title>3D BH<sub>3</sub> molecule</title> | ||
| + | <color>#ff82a8</color> | ||
| + | <size>200</size> | ||
| + | <uploadedFileContents>KATW NH3BH3 OPT.LOG</uploadedFileContents> | ||
| + | </jmolApplet></jmol> | ||
| + | |||
| + | ====Key bond distances and angles==== | ||
| + | {| class="wikitable" | ||
| + | |r(N-H) bond distance|| 1.02Â | ||
| + | |- | ||
| + | |r(B-H) bond distance|| 1.21Â | ||
| + | |- | ||
| + | | θ(H-N-H) bond angle|| 108° | ||
| + | |- | ||
| + | | θ(H-B-H) bond angle|| 114° | ||
| + | |- | ||
| + | | θ(H-B-N) bond angle|| 105° | ||
| + | |- | ||
| + | | θ(H-N-B) bond angle|| 111° | ||
| + | |} | ||
| + | |||
| + | |||
| + | ===File history=== | ||
| + | [[Media:KATW NH3BH3 OPT.LOG]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ===NH<sub>3</sub>BH<sub>3</sub> frequencies=== | ||
| + | <pre> | ||
| + | Low frequencies --- -2.7658 -2.2718 -0.0013 -0.0013 -0.0010 2.9530 | ||
| + | Low frequencies --- 263.4333 632.9750 638.4461 | ||
| + | </pre> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ==Lab1 Marking== | ||
| + | It's good that you have a working wiki. However, data in your item table don't match the values in the log file, and you have missed to include the torsion angle. If you have any queries, please contact Prof. Hunt. | ||
| + | |||
== NH<sub>3</sub> molecule == | == NH<sub>3</sub> molecule == | ||
===Calculation data=== | ===Calculation data=== | ||
| Line 10: | Line 485: | ||
|basis set|| 6-31G(d,p) | |basis set|| 6-31G(d,p) | ||
|- | |- | ||
| − | |final energy|| -56. | + | |final energy|| -56.557769au |
|- | |- | ||
| − | |RMS gradient||1.53e- | + | |RMS gradient||1.53e-07au/bohr |
|- | |- | ||
|point group|| C3v | |point group|| C3v | ||
| Line 26: | Line 501: | ||
</pre> | </pre> | ||
| − | ===Key bond distances and angles=== | + | ===Optimised molecule image=== |
| + | [[File:KATW nh3opt1finalimage.png|150px]] | ||
| + | <jmol><jmolApplet> | ||
| + | <title>3D NH<sub>3</sub> molecule</title> | ||
| + | <color>#ff82a8</color> | ||
| + | <size>200</size> | ||
| + | <uploadedFileContents>KATW307 NH3OPT1.LOG</uploadedFileContents> | ||
| + | </jmolApplet></jmol> | ||
| + | ====Key bond distances and angles==== | ||
{| class="wikitable" | {| class="wikitable" | ||
|r(N-H) bond distance|| 1.02Â | |r(N-H) bond distance|| 1.02Â | ||
| Line 32: | Line 515: | ||
| θ(H-N-H) bond angle|| 106° | | θ(H-N-H) bond angle|| 106° | ||
|} | |} | ||
| − | |||
| − | |||
| − | |||
===File history=== | ===File history=== | ||
[[Media:KATW307 NH3OPT1.LOG]] | [[Media:KATW307 NH3OPT1.LOG]] | ||
| − | + | ===NH<sub>3</sub> IR spectrum=== | |
| − | + | [[File:Katw NH3 irphoto.PNG]] | |
| − | |||
| − | |||
| − | |||
| − | |||
| + | ===NH<sub>3</sub> frequencies=== | ||
<pre> | <pre> | ||
Low frequencies --- -5.6864 -3.6131 -3.6124 0.0022 0.0047 0.0163 | Low frequencies --- -5.6864 -3.6131 -3.6124 0.0022 0.0047 0.0163 | ||
| Line 52: | Line 529: | ||
{| class="wikitable" | {| class="wikitable" | ||
| − | |mode||1||2||3||4||5 | + | |mode||1||2||3||4||5||6 |
|- | |- | ||
|wavenumber (cm<sup>-1</sup>)||1089||1694||1694||3461||3590||3590 | |wavenumber (cm<sup>-1</sup>)||1089||1694||1694||3461||3590||3590 | ||
| Line 62: | Line 539: | ||
===NH<sub>3</sub> charges=== | ===NH<sub>3</sub> charges=== | ||
| − | [[File:Katw nh3 | + | [[File:Katw nh3 chargemapfinal.PNG|500px]] |
{| class="wikitable" | {| class="wikitable" | ||
|Atom||Charge (e) | |Atom||Charge (e) | ||
| Line 71: | Line 548: | ||
|} | |} | ||
| − | |||
| − | |||
| − | == | + | |
| + | == N<sub>2</sub>F<sub>2</sub> molecule== | ||
===Calculation data=== | ===Calculation data=== | ||
| Line 86: | Line 562: | ||
|basis set|| 6-31G(d,p) | |basis set|| 6-31G(d,p) | ||
|- | |- | ||
| − | |final energy|| -309. | + | |final energy|| -309.0124au |
|- | |- | ||
| − | |RMS gradient||1.218e- | + | |RMS gradient||1.218e-06au/bohr |
|- | |- | ||
|point group|| C2v | |point group|| C2v | ||
| Line 102: | Line 578: | ||
</pre> | </pre> | ||
| − | ===Key bond distances and angles=== | + | ===Optimised molecule image=== |
| + | [[File:KATW n2f2opt1image.PNG|150px]] | ||
| + | <jmol><jmolApplet> | ||
| + | <title>3D NH<sub>3</sub> molecule</title> | ||
| + | <color>#ff82a8</color> | ||
| + | <size>200</size> | ||
| + | <uploadedFileContents>KATW307 N2F2OPT1.log</uploadedFileContents> | ||
| + | </jmolApplet></jmol> | ||
| + | ====Key bond distances and angles==== | ||
{| class="wikitable" | {| class="wikitable" | ||
|r(N-N) bond distance|| 1.22Â | |r(N-N) bond distance|| 1.22Â | ||
| Line 110: | Line 594: | ||
| θ(N-N-F) bond angle|| 114° | | θ(N-N-F) bond angle|| 114° | ||
|} | |} | ||
| − | |||
| − | |||
| − | |||
===File history=== | ===File history=== | ||
[[Media:KATW307 N2F2OPT1.log]] | [[Media:KATW307 N2F2OPT1.log]] | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| + | ===N<sub>2</sub>F<sub>2</sub> IR spectrum=== | ||
| + | [[File:KATW n2f2IR.PNG]] | ||
| + | |||
| + | ===N<sub>2</sub>F<sub>2</sub> frequencies=== | ||
<pre> | <pre> | ||
| − | Low frequencies --- - | + | Low frequencies --- -0.0011 0.0008 0.0019 3.4929 4.3715 5.1621 |
| − | Low frequencies --- | + | Low frequencies --- 347.8732 561.2460 771.6039 |
</pre> | </pre> | ||
{| class="wikitable" | {| class="wikitable" | ||
| − | |mode||1||2||3||4||5 | + | |mode||1||2||3||4||5||6 |
|- | |- | ||
| − | |wavenumber (cm<sup>-1</sup>)|| | + | |wavenumber (cm<sup>-1</sup>)||348||561||772||949||987||1637 |
|- | |- | ||
| − | |symmetry||A1|| | + | |symmetry||A1||A2||B2||A1||B2||A1 |
|- | |- | ||
| − | |intensity|| | + | |intensity (arbitrary units)||1||0||75||75||81||21 |
|} | |} | ||
| − | === | + | ===N<sub>2</sub>F<sub>2</sub> charges=== |
| − | [[File: | + | [[File:KATW n2f2opt1chargemap.PNG|500px]] |
{| class="wikitable" | {| class="wikitable" | ||
|Atom||Charge (e) | |Atom||Charge (e) | ||
|- | |- | ||
| − | |N|| | + | |N<sub>2</sub>|| 0.22 |
|- | |- | ||
| − | | | + | |F<sub>2</sub>||-0.22 |
|} | |} | ||
| − | === | + | ==Questions and answers== |
| − | [[File: | + | |
| + | <b>1. The molecule from the log file does not have bonds between the F and N atoms, what is going on here?</b> | ||
| + | |||
| + | The program running the optimization works under an assumption of what the length of the F and N atoms should be, and since the optimization put their bond length as longer than this assumed value, the program did not include a bond there. As the atoms get closer the program will include a bond between the atoms. | ||
| + | |||
| + | <b>2. For your IR analysis how many vibrations are expected from the 3N-6 rule?</b> | ||
| + | |||
| + | 3N-6 where N is the number of atoms would give N=4 and 3(4)-6= 6 vibrational modes, this is consistent with the number of vibrational modes obtained in the optimization as well. | ||
| + | |||
| + | <b>3. Why are there only 4 peaks in the IR spectrum?</b> | ||
| + | |||
| + | The A1 vibrational mode at 348 cm<sup>-1</sup> and the A2 vibrational mode at 561 cm<sup>-1</sup> both have an intensity of close to zero and were not detected by the IR | ||
| + | |||
| + | <b>4. For your IR analysis which vibration is the asymmetric N-F stretch?</b> | ||
| + | |||
| + | Vibrational mode 3 | ||
| + | |||
| + | <b>5. For your IR analysis what is the nature of the highest energy vibration?</b> | ||
| + | |||
| + | Symmetrical N-N bond stretching, vibrational mode 6 | ||
| + | |||
| + | <b>6. Which MOs are core orbital MOs? </b> | ||
| + | |||
| + | The 1-4 molecular orbitals which are only tied to their respective nuclei and not involved in bonding | ||
| + | |||
| + | <b>7. Provide a picture of MO 9 and beside it the LCAO diagram</b> | ||
| + | |||
| + | [[File:KATW n2f2mol9.PNG]] [[File:LCAO KATW.png|450px]] | ||
Latest revision as of 05:56, 15 May 2026
Contents
- 1 to do
- 2 1-methyl-imidazolium chloride
- 2.1 Calculation data
- 2.2 Item table
- 2.3 File history
- 2.4 1-methyl-imidazolium chloride A frequencies
- 2.5 Calculation data
- 2.6 Item table
- 2.7 File history
- 2.8 1-methyl-imidazolium chloride B frequencies
- 2.9 Calculation data
- 2.10 Item table
- 2.11 File history
- 2.12 1-methyl-imidazolium chloride C frequencies
- 2.13 1-methyl-imidazolium chloride A scan of total energy
- 3 Me3NH Cl molecule
- 4 Relative energies
- 5 BH3 molecule
- 6 NH3BH3 molecule
- 7 Lab1 Marking
- 8 NH3 molecule
- 9 N2F2 molecule
- 10 Questions and answers
to do
- redo some of the accuracies
tabulate and compare all the H---Cl distances for all ion-pairs. how do the H-bonds of Me3NH and HMim compare? how do the H-bonds of the N-H and C-H H-bonds compare? are these distances representative of a H-bond? Hint compare your distances to the sum of the van der waals radii of H and Cl. will the ionic nature of the ions effect a distance based assessment of H-bonding? provide a table and comparison of the association energy for a-c make this explicit, so give the energy of each component in atomic units, compute the deltaE in au and then convert to kJ/mol provide the relative energy of the two isomers (a) and (b), ask if you are not sure what this is. rationalise why one conformer is less stable than the other. discuss the dissociation energy of (c) relative to (a) and (b). What does the comparison tell us about the H-bonding? scan the N-H coordinate towards Cl of (a) take your optimised (a)structure, swap atoms to make N=1 and H=2, set the N-H distance to 0.8 and the N-Cl distance to 3.2, run the scan. This will take some time, start it early! provide a snapshot from gaussview of the "raw" HMim-Cl PES plot provide a chemical diagram of the two protonation states for HMim-Cl graph the MeNH-Cl and HMim-Cl scans on a single PES plot discuss your HMim-Cl PES plot, compare and contrast your results for the MeNH-Cl and HMim-Cl PES.
1-methyl-imidazolium chloride
Calculation data
| name of submitted log file | KATW_RINGA_OPTFREQ2.log |
| molecule | 1-methyl-imidazolium chloride A |
| method | RB3LYP |
| basis set | 3-21G |
| final energy | -722.68731au |
| RMS gradient | 5.198e-06au/bohr |
| point group | C1 |
Item table
Optimizing item table
Item Value Threshold Converged? Maximum Force 0.000011 0.000450 YES RMS Force 0.000003 0.000300 YES Maximum Displacement 0.001137 0.001800 YES RMS Displacement 0.000246 0.001200 YES
Frequency item table
Item Value Threshold Converged? Maximum Force 0.000011 0.000450 YES RMS Force 0.000003 0.000300 YES Maximum Displacement 0.001332 0.001800 YES RMS Displacement 0.000305 0.001200 YES
Key bond distances and angles
- add another decimal point to each
| r(H-Cl) bond distance | 1.719Â |
File history
1-methyl-imidazolium chloride A frequencies
Low frequencies --- -5.2962 -2.8134 -0.0015 0.0024 0.0026 2.7809 Low frequencies --- 36.1361 64.4953 80.8261
The initial frequency scan resulted in a negative frequency vibration from the methyl group. The methyl group was manually rotated and the scan run again with reading internal force constants to yield better data.
Calculation data
| name of submitted log file | KATW RINGB OPTFREQ2.LOG |
| molecule | 1-methyl-imidazolium chloride B |
| method | RB3LYP |
| basis set | 3-21G |
| final energy | -722.6662au |
| RMS gradient | 2.5631e-05au/bohr |
| point group | C1 |
Item table
Optimizing item table
Item Value Threshold Converged? Maximum Force 0.000088 0.000450 YES RMS Force 0.000020 0.000300 YES Maximum Displacement 0.000693 0.001800 YES RMS Displacement 0.000163 0.001200 YES
Frequency item table
Item Value Threshold Converged? Maximum Force 0.000088 0.000450 YES RMS Force 0.000020 0.000300 YES Maximum Displacement 0.001343 0.001800 YES RMS Displacement 0.000321 0.001200 YES
Key bond distances and angles
- add another decimal point to each
| r(CH-Cl) bond distance | 2.134Â |
| r(CH2H-Cl) bond distance | 2.277Â |
File history
1-methyl-imidazolium chloride B frequencies
Low frequencies --- -6.2380 -3.1641 0.0011 0.0033 0.0040 1.3698 Low frequencies --- 45.3380 162.1734 198.8517
Calculation data
| name of submitted log file | KATW_RINGC_OPTFREQ.log |
| molecule | 1-methyl-imidazolium chloride C |
| method | RB3LYP |
| basis set | 3-21G |
| final energy | -761.77953au |
| RMS gradient | 8.305e-06au/bohr |
| point group | C1 |
Item table
Optimizing item table
Item Value Threshold Converged? Maximum Force 0.000027 0.000450 YES RMS Force 0.000005 0.000300 YES Maximum Displacement 0.000555 0.001800 YES RMS Displacement 0.000154 0.001200 YES
Frequency item table
Item Value Threshold Converged? Maximum Force 0.000027 0.000450 YES RMS Force 0.000005 0.000300 YES Maximum Displacement 0.000581 0.001800 YES RMS Displacement 0.000167 0.001200 YES
Key bond distances and angles
| r(H-Cl) bond distance | 2.030Â |
File history
1-methyl-imidazolium chloride C frequencies
Low frequencies --- -3.7052 -3.0699 -0.0042 -0.0027 -0.0019 2.2969 Low frequencies --- 52.1621 102.5835 107.1578
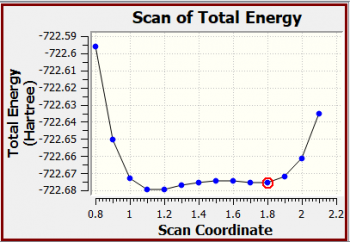
1-methyl-imidazolium chloride A scan of total energy
Scan of Total Energy
X-Axis: Scan Coordinate
Y-Axis: Total Energy (Hartree)
X Y
0.8000000000 -722.5958115700
0.9000000000 -722.6503976760
1.0000000000 -722.6727649280
1.1000000000 -722.6793731750
1.2000000000 -722.6792905430
1.3000000000 -722.6772128260
1.4000000000 -722.6753243350
1.5000000000 -722.6744432350
1.6000000000 -722.6746287060
1.7000000000 -722.6753547910
1.8000000000 -722.6753495180
1.9000000000 -722.6721376400
2.0000000000 -722.6613018340
2.1000000000 -722.6354480070
Me3NH Cl molecule
Calculation data
| name of submitted log file | KATW_Me3NH-Clactual.log |
| molecule | Me3NH Cl |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -635.31113au |
| RMS gradient | 4.0214e-05au/bohr |
| point group | C1 |
The above point group was not set to be C3v, which it should be, so the program assigned it C1.
Item table
Optimizing item table
Item Value Threshold Converged? Maximum Force 0.000126 0.000450 YES RMS Force 0.000032 0.000300 YES Maximum Displacement 0.000835 0.001800 YES RMS Displacement 0.000219 0.001200 YES
Frequency item table
Item Value Threshold Converged? Maximum Force 0.000126 0.000450 YES RMS Force 0.000032 0.000300 YES Maximum Displacement 0.001285 0.001800 YES RMS Displacement 0.000457 0.001200 YES
Optimised molecule image
3D Me3NH Cl molecule |
Key bond distances and angles
- add another decimal point to each
| r(N-C) bond distance | 1.48Â |
| r(N-H) bond distance | 0.80Â |
| r(N-Cl) bond distance | 3.20Â |
| r(C-H) bond distance | 1.09Â |
| θ(C-N-C) bond angle | 112° |
| θ(C-N-H) bond angle | 106° |
| θ(H-C-H) bond angle | 110° |
File history
NH3BH3 frequencies
Low frequencies --- -3.0077 -0.5001 -0.0059 -0.0053 -0.0047 6.2314 Low frequencies --- 81.5384 81.6770 183.6441
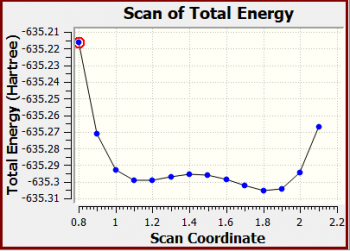
NH3BH3 scan of total energy
Snapshot:
X-Axis: Scan Coordinate
Y-Axis: Total Energy (Hartree)
X Y
0.8000000000 -635.2162861320
0.9000000000 -635.2709452560
1.0000000000 -635.2930688430
1.1000000000 -635.2992906270
1.2000000000 -635.2988745370
1.3000000000 -635.2968157070
1.4000000000 -635.2956086330
1.5000000000 -635.2962523740
1.6000000000 -635.2987711990
1.7000000000 -635.3023546920
1.8000000000 -635.3052254510
1.9000000000 -635.3042681400
2.0000000000 -635.2943300920
2.1000000000 -635.2670421330
Scan of the potential energy surface original graph:
Relative energies
| molecule | Energy (au) | Energy (kJ/mol) | 3 | 4 | 5 | 6 |
| NH3 | -56.557769 | -148492 | 1213 | 2583 | 2716 | 2716 |
| BH3 | -26.615324 | -69879 | E | A1 | E | E |
| NH3BH3 | -83.2246894 | -218506 | 14 | 0 | 126 | 126 |
E(NH3BH3)=
BH3 molecule
Calculation data
| name of submitted log file | BH3_optimization_KATW.log |
| molecule | BH3 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -26.615324au |
| RMS gradient | 2.114e-06au/bohr |
| point group | D3H |
Item table
Item Value Threshold Converged? Maximum Force 0.000004 0.000015 YES RMS Force 0.000003 0.000010 YES Maximum Displacement 0.000017 0.000060 YES RMS Displacement 0.000011 0.000040 YES
Optimised molecule image
3D BH molecule |
Key bond distances and angles
| r(B-H) bond distance | 1.19Â |
| θ(H-B-H) bond angle | 120° |
File history
Media:BH3 OPTIMIZATION KATW.LOG
BH3 IR spectrum
BH3 frequencies
Low frequencies --- -11.6940 -11.6861 -6.5543 -0.0006 0.0280 0.4289 Low frequencies --- 1162.9745 1213.1390 1213.1392
| mode | 1 | 2 | 3 | 4 | 5 | 6 |
| wavenumber (cm-1) | 1163 | 1213 | 1213 | 2583 | 2716 | 2716 |
| symmetry | A2 | E | E | A1 | E | E |
| intensity | 93 | 14 | 14 | 0 | 126 | 126 |
NH3BH3 molecule
Calculation data
| name of submitted log file | KATW_NH3BH3_opt.log |
| molecule | NH3BH3 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -83.2246894au |
| RMS gradient | 1.264e-06au/bohr |
| point group | C1 |
Item table
Item Value Threshold Converged? Maximum Force 0.000002 0.000015 YES RMS Force 0.000001 0.000010 YES Maximum Displacement 0.000016 0.000060 YES RMS Displacement 0.000007 0.000040 YES
Optimised molecule image
3D BH molecule |
Key bond distances and angles
| r(N-H) bond distance | 1.02Â |
| r(B-H) bond distance | 1.21Â |
| θ(H-N-H) bond angle | 108° |
| θ(H-B-H) bond angle | 114° |
| θ(H-B-N) bond angle | 105° |
| θ(H-N-B) bond angle | 111° |
File history
NH3BH3 frequencies
Low frequencies --- -2.7658 -2.2718 -0.0013 -0.0013 -0.0010 2.9530 Low frequencies --- 263.4333 632.9750 638.4461
Lab1 Marking
It's good that you have a working wiki. However, data in your item table don't match the values in the log file, and you have missed to include the torsion angle. If you have any queries, please contact Prof. Hunt.
NH3 molecule
Calculation data
| name of submitted log file | KATW307_nh3opt1.log |
| molecule | NH3 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -56.557769au |
| RMS gradient | 1.53e-07au/bohr |
| point group | C3v |
Item table
Item Value Threshold Converged? Maximum Force 0.000002 0.000015 YES RMS Force 0.000001 0.000010 YES Maximum Displacement 0.000010 0.000060 YES RMS Displacement 0.000006 0.000040 YES
Optimised molecule image
3D NH molecule |
Key bond distances and angles
| r(N-H) bond distance | 1.02Â |
| θ(H-N-H) bond angle | 106° |
File history
NH3 IR spectrum
NH3 frequencies
Low frequencies --- -5.6864 -3.6131 -3.6124 0.0022 0.0047 0.0163 Low frequencies --- 1089.3674 1693.9284 1693.9284
| mode | 1 | 2 | 3 | 4 | 5 | 6 |
| wavenumber (cm-1) | 1089 | 1694 | 1694 | 3461 | 3590 | 3590 |
| symmetry | A1 | E | E | A1 | E | E |
| intensity | 145 | 14 | 14 | 1 | 0 | 0 |
NH3 charges
| Atom | Charge (e) |
| N | -1.13 |
| H3 | 0.38 |
N2F2 molecule
Calculation data
| name of submitted log file | KATW307_n2f2opt1.log |
| molecule | N2F2 |
| method | RB3LYP |
| basis set | 6-31G(d,p) |
| final energy | -309.0124au |
| RMS gradient | 1.218e-06au/bohr |
| point group | C2v |
Item table
Item Value Threshold Converged? Maximum Force 0.000000 0.000015 YES RMS Force 0.000000 0.000010 YES Maximum Displacement 0.000003 0.000060 YES RMS Displacement 0.000001 0.000040 YES
Optimised molecule image
3D NH molecule |
Key bond distances and angles
| r(N-N) bond distance | 1.22Â |
| r(N-F) bond distance | 1.39Â |
| θ(N-N-F) bond angle | 114° |
File history
N2F2 IR spectrum
N2F2 frequencies
Low frequencies --- -0.0011 0.0008 0.0019 3.4929 4.3715 5.1621 Low frequencies --- 347.8732 561.2460 771.6039
| mode | 1 | 2 | 3 | 4 | 5 | 6 |
| wavenumber (cm-1) | 348 | 561 | 772 | 949 | 987 | 1637 |
| symmetry | A1 | A2 | B2 | A1 | B2 | A1 |
| intensity (arbitrary units) | 1 | 0 | 75 | 75 | 81 | 21 |
N2F2 charges
| Atom | Charge (e) |
| N2 | 0.22 |
| F2 | -0.22 |
Questions and answers
1. The molecule from the log file does not have bonds between the F and N atoms, what is going on here?
The program running the optimization works under an assumption of what the length of the F and N atoms should be, and since the optimization put their bond length as longer than this assumed value, the program did not include a bond there. As the atoms get closer the program will include a bond between the atoms.
2. For your IR analysis how many vibrations are expected from the 3N-6 rule?
3N-6 where N is the number of atoms would give N=4 and 3(4)-6= 6 vibrational modes, this is consistent with the number of vibrational modes obtained in the optimization as well.
3. Why are there only 4 peaks in the IR spectrum?
The A1 vibrational mode at 348 cm-1 and the A2 vibrational mode at 561 cm-1 both have an intensity of close to zero and were not detected by the IR
4. For your IR analysis which vibration is the asymmetric N-F stretch?
Vibrational mode 3
5. For your IR analysis what is the nature of the highest energy vibration?
Symmetrical N-N bond stretching, vibrational mode 6
6. Which MOs are core orbital MOs?
The 1-4 molecular orbitals which are only tied to their respective nuclei and not involved in bonding
7. Provide a picture of MO 9 and beside it the LCAO diagram